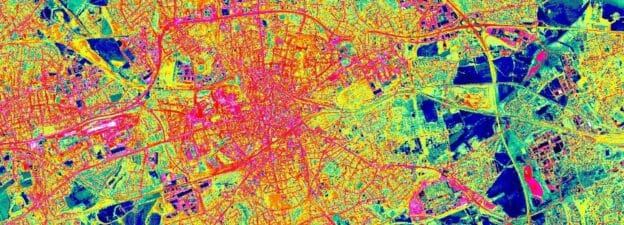
Urbanization is changing our planet. About half of the world’s population already lives in cities. By 2050 that proportion is expected to rise to two-thirds. The loss of vegetation and the sprawl of impervious surfaces that are connected to this process are the cause of many problems societies have to face in the future. On the social level cities in the southern hemisphere are experiencing an uptick in unregulated settling resulting in the sprawl of slums, leaving large amounts of people in dire situations cut off from the most basic needs of daily life.
Remote sensing can help to better understand those problems by capturing the changes in urban areas. Besides just visual observations, it also can provide quantification and more detailed information on the impacts of those changes, and thus provide decision-makers with valuable information.
The hands-on lessons in this course will teach you:
- How to map urban areas using high-resolution data in Google Earth Engine
- How to quantify urban heat islands in cities in Google Earth Engine
- How to visually delineate slum areas
- How to predict slum areas using an advanced classification algorithm in R for the city of Lagos in Nigeria
Urban Land Cover Mapping
Learning objectives of this lesson Understand the current challenges of urban monitoring Overview of data and methods of urban monitoring…
Mapping Slums
Learning objectives of this lesson Understand slum definition and development. Learn about approaches to mapping slums. Visually identify and map…
Note: you do not have to take the lessons in any particular order.
Related courses
If you are interested in more ‘Land in Focus’ courses, check out the availability below. The courses will be available in the future and will be released sequentially towards spring of 2022. These will provide you with theoretical knowledge and real-life case studies from disaster management, to land cover change analysis all the way to food security-related issues, that you can work with using the tools of remote sensing.
Funded by
Course Content
About Instructor